Modulation of the chromatin structure plays an important role in the regulation of transcription in eukaryotes. The nucleosome, made up of four core histone proteins (H2A, H2B, H3 and H4), is the primary building block of chromatin. The N-terminal tail of core histones undergoes different posttranslational modifications including acetylation, phosphorylation and methylation. These modifications occur in response to cell signal stimuli and have a direct effect on gene expression. In most species, the histone H2B is primarily acetylated at lysines 5, 12, 15 and 20. Histone H3 is primarily acetylated at lysines 9, 14, 18 and 23. Acetylation at lysine 9 appears to have a dominant role in histone deposition and chromatin assembly in some organisms. Phosphorylation at Ser10 of histone H3 is tightly correlated with chromosome condensation during both mitosis and meiosis.
Function:
Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. H3 is deposited into chromatin exclusively through a DNA replication-coupled pathway that can be associated with either DNA duplication or DNA repair synthesis during meiotic homologous recombination.
Subunit:
The nucleosome is a histone octamer containing two molecules each of H2A, H2B, H3 and H4 assembled in one H3-H4 heterotetramer and two H2A-H2B heterodimers. The octamer wraps approximately 147 bp of DNA. Interacts with GCN5, whereby H3S10ph increases histone-protein interactions. Interacts with PDD1 and PDD3.
Subcellular Location:
Nucleus. Chromosome. Note=Localizes to both the large, transcriptionally active, somatic macronucleus (MAC) and the small, transcriptionally inert, germ line micronucleus (MIC).
Post-translational modifications:
Phosphorylated to form H3S10ph. H3S10ph promotes subsequent H3K14ac formation by GCN5. H3S10ph is only found in the mitotically dividing MIC, but not in the amitotically dividing MAC. H3S10ph is correlated with chromosome condensation during mitotic or meiotic micronuclear divisions.
Acetylation of histone H3 leads to transcriptional activation. H3K14ac formation by GCN5 is promoted by H3S10ph. H3K9acK14ac is the preferred acetylated form of newly synthesized H3. Acetylation occurs almost exclusively in the MAC.
Methylated to form H3K4me. H3K4me is only found in the transcriptionally active MAC. Methylated to form H3K9me in developing MACs during conjugation, when genome-wide DNA elimination occurs. At this stage, H3K9me specifically occurs on DNA sequences being eliminated (IES), probably targeted by small scan RNAs (scnRNAs) bound to IES, and is required for efficient IES elimination. H3K9me is required for the interaction with the chromodomains of PDD1 and PDD3.
The full-length protein H3S (slow migrating) is converted to H3F (fast migrating) by proteolytic removal of the first 6 residues. H3F is unique to MIC, and processing seems to occur regularly each generation at a specific point in the cell cycle.
Similarity:
Belongs to the histone H3 family.
SWISS:
P68431
Gene ID:
8350
Database links:
Entrez Gene: 8350 Human
Entrez Gene: 8351 Human
Entrez Gene: 8352 Human
Entrez Gene: 8353 Human
Entrez
Gene: 8354 Human
Entrez Gene: 8355 Human
Entrez Gene: 8356 Human
Entrez Gene: 8357 Human
Entrez Gene: 8358 Human
Entrez Gene: 8968 Human
Entrez Gene: 260423 Mouse
Entrez Gene: 319148 Mouse
Entrez Gene: 319149 Mouse
Entrez Gene: 319150 Mouse
Entrez Gene: 319151 Mouse
Entrez Gene: 319152 Mouse
Entrez Gene: 319153 Mouse
Entrez Gene: 72198 Mouse
Entrez Gene: 97908 Mouse
Entrez Gene: 100364501 Rat
Entrez Gene: 100365669 Rat
Entrez Gene: 291159 Rat
Entrez Gene: 314977 Rat
Entrez Gene: 364716 Rat
Entrez Gene: 679950 Rat
Entrez Gene: 679994 Rat
Entrez Gene: 136511 Rat
Entrez Gene: 136599 Rat
Entrez Gene: 682330 Rat
Entrez Gene: 691496 Rat
SwissProt: P68431 Human
SwissProt: P84243 Human
SwissProt: Q16695 Human
SwissProt: Q6NXT2 Human
SwissProt: Q71DI3 Human
SwissProt: P68433 Mouse
SwissProt: P84228 Mouse
SwissProt: Q6LED0 Rat
| Picture |
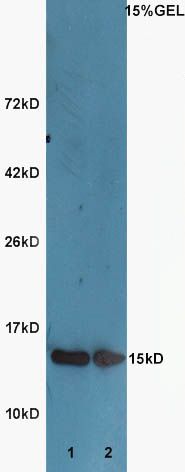
Sample:
Lane1:Huh7 Cell Lysate at 40 ug
Lane2:A549 Cell Lysate at 40 ug
Primary: Anti-Histone H3 (SL3776R) at 1:300 dilution;
Secondary: HRP conjugated Goat-Anti-Rabbit IgG(SL0295G-HRP) at 1: 5000 dilution;
Predicted band size :15 kD
Observed band size :15 kD
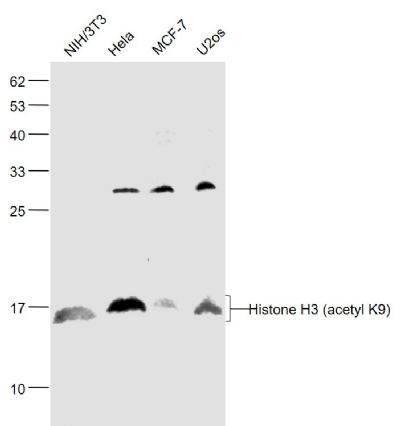
Sample:
NIH/3T3(Mouse) Cell Lysate at 30 ug
Hela(Human) Cell Lysate at 30 ug
MCF-7(Human) Cell Lysate at 30 ug
U2os(Human) Cell Lysate at 30 ug
Primary: Anti-Histone H3 (acetyl K9) (SL3776R) at 1/1000 dilution
Secondary: IRDye800CW Goat Anti-Rabbit IgG at 1/20000 dilution
Predicted band size: 15 kD
Observed band size: 15 kD
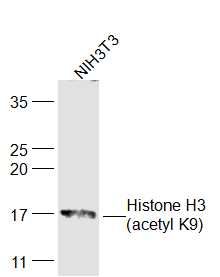
Sample:
NIH/3T3(Mouse) Cell Lysate at 30 ug
Primary: Anti-Histone H3 (acetyl K9) (SL3776R) at 1/300 dilution
Secondary: IRDye800CW Goat Anti-Rabbit IgG at 1/20000 dilution
Predicted band size: 15 kD
Observed band size: 17 kD
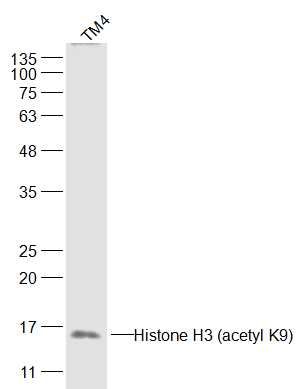
Sample:
TM4(Human) Cell Lysate at 30 ug
Primary: Anti-Histone H3 (acetyl K9) (SL3776R) at 1/300 dilution
Secondary: IRDye800CW Goat Anti-Rabbit IgG at 1/20000 dilution
Predicted band size: 15 kD
Observed band size: 15 kD
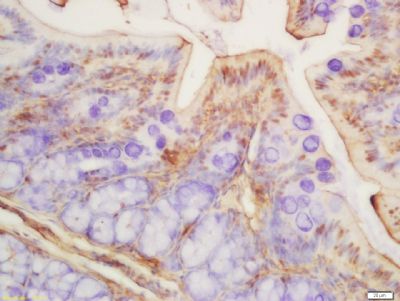
Tissue/cell: rat colon tissue; 4% Paraformaldehyde-fixed and paraffin-embedded;
Antigen retrieval: citrate buffer ( 0.01M, pH 6.0 ), Boiling bathing for 15min; Block endogenous peroxidase by 3% Hydrogen peroxide for 30min; Blocking buffer (normal goat serum,SLC0005) at 37℃ for 20 min;
Incubation: Anti-Histone H3 (acetyl K9) Polyclonal Antibody, Unconjugated(SL3776R) 1:200, overnight at 4°C, followed by conjugation to the secondary antibody(SP-0023) and DAB(SLC0010) staining
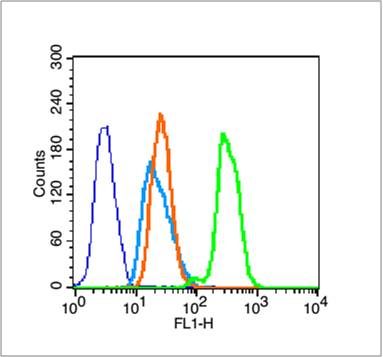
Blank control (blue line): Hela (fixed with 70% ethanol (Overnight at 4℃) and then permeabilized with 90% ice-cold methanol for 30 min on ice).
Primary Antibody (green line): Rabbit Anti-Histone H3 (acetyl K9) antibody (SL3776R),Dilution: 1μg /10^6 cells;
Isotype Control Antibody (orange line): Rabbit IgG .
Secondary Antibody (white blue line): Goat anti-rabbit IgG-FITC,Dilution: 1μg /test.
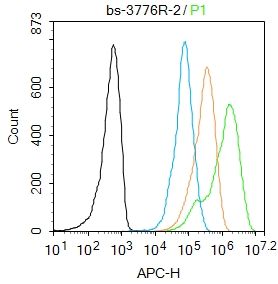
Blank control: Molt4.
Primary Antibody (green line): Rabbit Anti-Histone H3 (acetyl K9) antibody (SL3776R)
Dilution: 2μg /10^6 cells;
Isotype Control Antibody (orange line): Rabbit IgG .
Secondary Antibody : Goat anti-rabbit IgG-AF647
Dilution: 1μg /test.
Protocol
The cells were fixed with 4% PFA (10min at room temperature)and then permeabilized with 90% ice-cold methanol for 20 min at -20℃. The cells were then incubated in 5%BSA to block non-specific protein-protein interactions for 30 min at room temperature .Cells stained with Primary Antibody for 30 min at room temperature. The secondary antibody used for 40 min at room temperature. Acquisition of 20,000 events was performed.
|
|
|