This gene was identified based on homology to Pichia pastoris GSA7 and Saccharomyces cerevisiae APG7. In the yeast, the protein appears to be required for fusion of peroxisomal and vacuolar membranes. The protein shows homology to the ATP-binding and catalytic sites of the E1 ubiquitin activating enzymes. [provided by RefSeq, Jan 2009].
Function:
Functions as an E1 enzyme essential for multisubstrates such as ATG8-like proteins and ATG12. Forms intermediate conjugates with ATG8-like proteins (GABARAP, GABARAPL1, GABARAPL2 or MAP1LC3A). PE-conjugation to ATG8-like proteins is essential for autophagy. Also acts as an E1 enzyme for ATG12 conjugation to ATG5 and ATG3.
Subunit:
Homodimer. Interacts with ATG3 and ATG12. The complex, composed of ATG3 and ATG7, plays a role in the conjugation of ATG12 to ATG5.
Subcellular Location:
Cytoplasm (Probable).
Tissue Specificity:
Widely expressed, especially in kidney, liver, lymph nodes and bone marrow.
Similarity:
Belongs to the ATG7 family.
SWISS:
O95352
Gene ID:
10533
Database links:
Entrez Gene: 10533 Human
Entrez Gene: 415961 Chicken
Entrez Gene: 74244 Mouse
Entrez Gene: 312647 Rat
Omim: 608760 Human
SwissProt: Q5ZKY2 Chicken
SwissProt: O95352 Human
SwissProt: Q9D906 Mouse
SwissProt: Q641Y5 Rat
Unigene: 38032 Human
Unigene: 275332 Mouse
Unigene: 162765 Rat
| Picture |
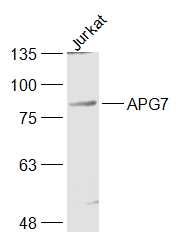
Sample:
Jurkat(Human) Cell Lysate at 30 ug
Primary: Anti-APG7 (SL2432R) at 1/1000 dilution
Secondary: IRDye800CW Goat Anti-Rabbit IgG at 1/20000 dilution
Predicted band size: 75 kD
Observed band size: 85 kD
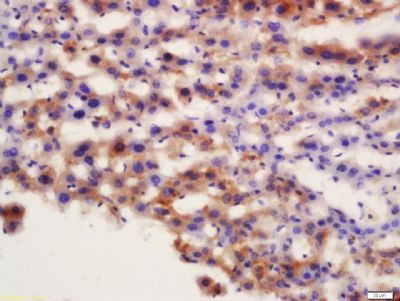
Tissue/cell: mouse liver tissue; 4% Paraformaldehyde-fixed and paraffin-embedded;
Antigen retrieval: citrate buffer ( 0.01M, pH 6.0 ), Boiling bathing for 15min; Block endogenous peroxidase by 3% Hydrogen peroxide for 30min; Blocking buffer (normal goat serum,SLC0005) at 37℃ for 20 min;
Incubation: Anti-APG7 Polyclonal Antibody, Unconjugated(SL2432R) 1:200, overnight at 4°C, followed by conjugation to the secondary antibody(SP-0023) and DAB(SLC0010) staining
|
|
|